Dr. Subhendu Kumar, Director – Research Operations, Integrated Drug Discovery, Syngene, moderated a panel discussion at the Targeted Protein Degradation & PROTAC Virtual Symposium, 2021, recently. The discussion covered some key topics related to the development of degraders from a Medicinal Chemistry perspective. Pre-eminent scientists in the field of protein degradation, from both industry and academia, participated in the discussion. Summarized below are some fascinating insights which emerged from this discussion:
Key considerations for developing protein degraders
Although the scientific community has gained significant expertise with degrading several targets, we are still a fair bit away from saying that we can degrade any protein. One of the key challenges is identifying the protein of interest (POI) for intractable targets, regardless of the mechanism of modulation. Several target classes have been explored recently including transmembrane proteins. Key considerations for the effectiveness of the approach would be turnover rates and expression levels of targets, among other things. Structural considerations such as the presence of lysines for ubiquitination, as well as their orientation, will help prioritize targets. Challenges for the TPD platform from a potential therapeutic modality would be to ensure oral-bioavailability and overcoming the blood-brain barrier for being effective against CNS indications.
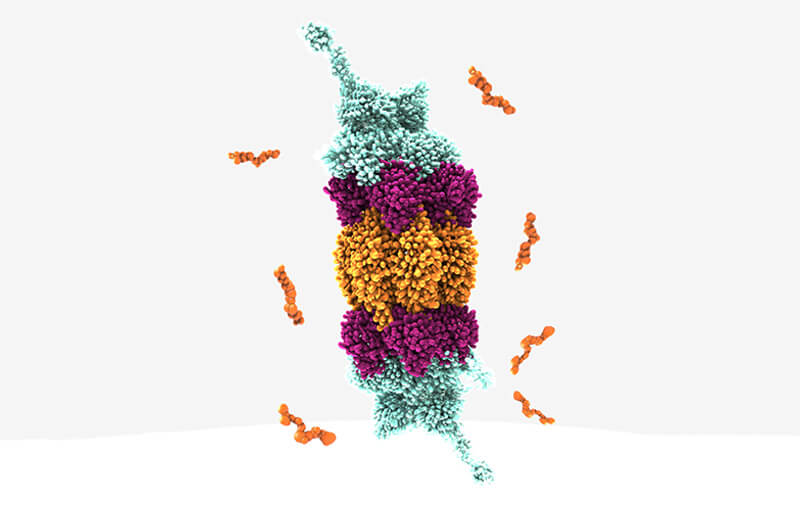
Structural biology of degradation
Although cryogenic electron microscopy has opened some new avenues for structural biology over crystallization, it is not about one technique but a combination of different available tools– biophysics, structural biology or crystallization. The actual combination will depend on the targets themselves. Degradation also necessitates being more open and broader with respect to molecule design, even if you have identified a crystal structure.
When to consider physicochemical properties — At the beginning of a program or during lead optimization
One key question for scientists in the field is when to consider physicochemical properties –at the beginning of a program or during lead optimization. Presently, we are at a stage where running experiments and generating data in this field, is of paramount importance. Property-based design can, however, help avoid the synthesis of compounds that are unlikely to become a viable therapeutic agent.
Future of Molecular glues and application in CNS
Innovative assay systems are needed to identify viable molecular glues. They are more promising for Central Nervous Systems (CNS) applications, with the possibility of getting you to a point where you can design compounds that can go past the blood brain barrier. That remains the key challenge – optimizing the degraders for brain penetration. As for how to screen for start points for molecular glues, two approaches are suggested:
1) Start with existing E3 ligases and ligands and build libraries
2) Phenotypic cell-based approach looking for degradation of specific targets followed by deconvolution exercise.
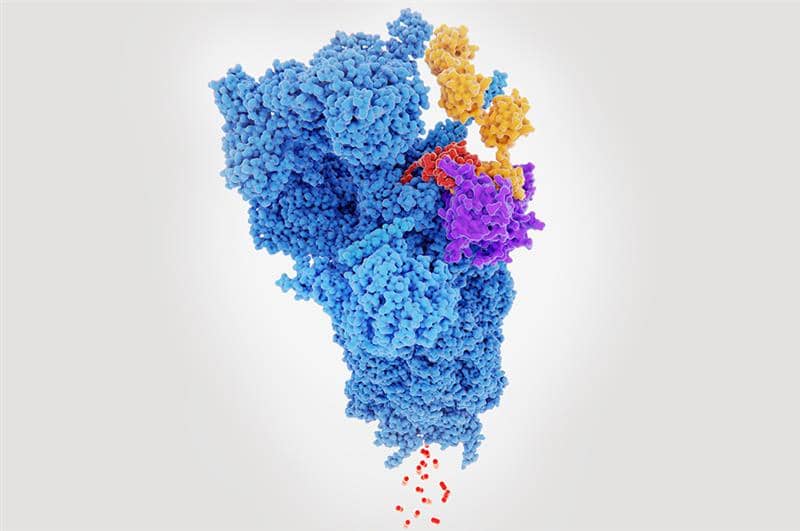
Key considerations in PROTAC design
Mechanism of relevance is the key consideration in PROTAC design i.e. stabilizing ternary complexes to get better degraders, and kinetic pathways vs thermodynamic equilibrium.
One key question is whether these should be considered while designing PROTACs if it is more a case of retrospective analysis to understand the mechanism. One approach is to build the assays that can identify the mechanism that is prevalent – stability vs. dynamism. After that, try and characterize as soon as you have an active compound, and collect the data to understand the rules and applicability across degraders.
Picking the right E3 ligase for your target
Since there are limited ligases currently, an experimental approach is viable to identify a suitable one. A bioinformatics approach evaluating all 600 and trying to narrow down is possible. However, there are limited E3 ligases with ligands discovered as of now – the list gets cut down quickly. Expression profile against different tissues can also be used to narrow down the list. Over the next few years, a commonly accepted panel of optimal ligases is likely to emerge.
To know about Syngene’s PROTAC capabilities, please click here